Inhibition of protein expression by the interaction of G-Quadruplex and RHAU peptide in E. coli
- Center for Bioscience and Biotechnology, University of Science, National University-HCMC, Vietnam
- Institue of Tropical Biology, VAST
- aculty of Biotechnology, Ho Chi Minh City Open University, Vietnam
Abstract
Introduction: G-quadurplex (G4) formation plays a role in many biological processes such as replication, transcription, translation and telomeric maintenance. Stabilization of G4 structure by peptide has recently emerged as a potential approach in the regulation of protein expression. In this study, we reported on regulation of cyan fluorescent protein (CFP) expression by the interaction of G4 and RNA helicase associated with AU-rich elements (RHAU) peptide in E. coli.
Methods: A sequence of TTGGGTGGGTGGGTGGGT (formed into G4 structure) was genetically applied to cfp gene as a reporter gene (g4-cfp). Both g4-cfp and DHX36 (or ΔDHX36) genes were cloned to pET-Duet1 vector that allowed to simultaneously express both G4-CFP protein and RHAU (or ΔRHAU) protein under IPTG inducer.
Results: The level of G4-CFP expression in the presence of RHAU (pD64) was around 2-fold and 3-fold lower than that of G4-CFP expression in the presence of ΔRHAU (pD65) and G4-CFP expression alone (pD21), respectively.
Conclusion: RHAU might selectively bind G4 structure of mRNA of G4-CFP, resulting in inhibition of G4-CFP expression in E. coli. The G4 and RHAU peptide interaction would provide a promising approach for inhibition of gene expression in many biomedical applications.
INTRODUCTION
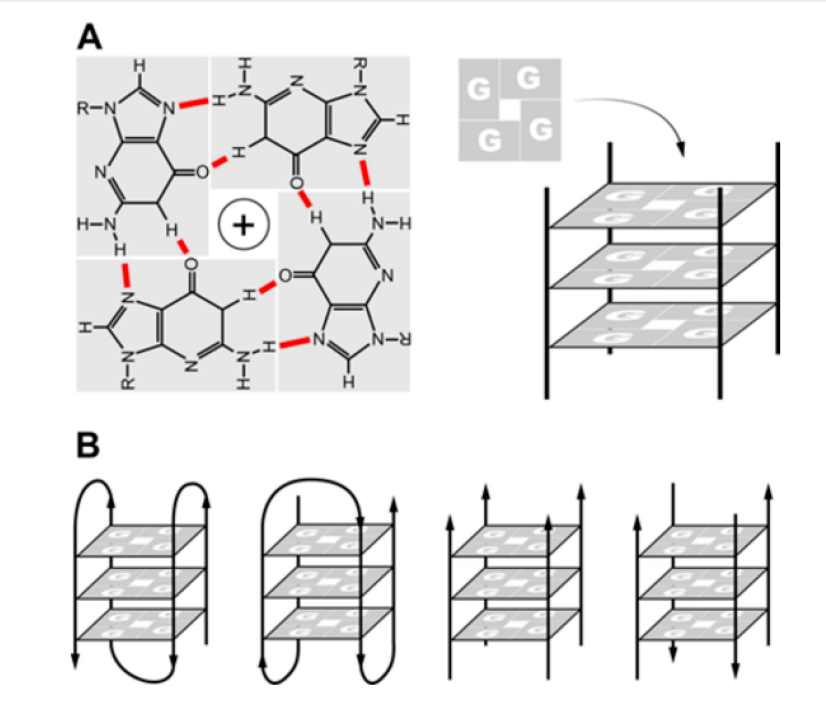
G-quadruplexes (G4s) are G-rich sequences that can fold into four single-stranded DNA or RNA structures by hoogsteen hydrogen interaction1 (Figure 1). G4 structures can be parallel or nonparallel topology2. In the cell, the formation of G4 structure plays a crucial role in many biological processes such as replication, transcription, translation and telomeric maintenance3. For instance, the formation of G4 in the untranslated region (UTR) that significantly affected the level expression of protein6, 5, 4. Formation of G4 structure at the telomeric region that can prevent binding of telomerase to its DNA target leads to inhibition of telomeric elongation9, 8, 7.Therefore, the formation of G4s in DNA or RNA is considered to be a new molecular target for molecules in cancer therapeutics12, 11, 10.
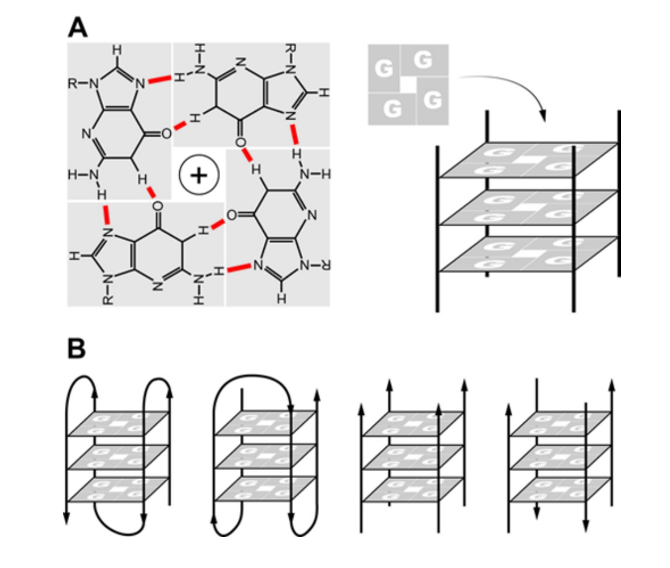
A) G4 structure is formed in the presence of cation K+ or Na+. B) G4s with different topologies: parallel and non-parallel
Specific recognition and stabilization of G4 by peptides have emerged as a potential approach for regulation of many biological processes due to the following advantages such as (i) peptides are easier to design and synthesize than recombinant proteins, (ii) peptides can mimic the interaction of G4 and protein. Recently, specific recognition of parallel G4 by RHAU has been reported13. The full length of RHAU protein consisting of 1008 aa can bind parallel G4 and unwind G4 structure in the presence of ATP in the cells. However, only N-region of RHAU peptide (without helicase domain) can specifically bind and stabilize parallel G4 structure13. Insight into the structure of G4 and RHAU peptide showed the RHAU covers a terminal guanine base and binds the parallel G4 using 3 anchor point electrostatic interactions between negative charge of phosphate groups and 3 positive charges of amino acids14. The studies of the interaction of G4 and RHAU peptide for biological applications have also been reported. Incorporating RHAU peptide to CFP allowed to generate the fluorescent probe which can visualize and distinguish G4 topologies15. Development of the novel ribonuclease by fusing RHAU with RNase H catalytic domain that can selectively bind G4 and cleave RNA in RNA:DNA hybrid at the specific manner16. However, the application of stabilization of G4 by RHAU peptide in inhibition of protein expression has not been reported yet.
Herein, we study inhibition of CFP expression by the interaction between G4 and RHAU peptide (140 aa) in E. coli (Figure 2). Two genes: g4-cfp (consisting G4 structure) and DHX36 were genetically cloned in the pET-Duet1 vector that allows expressing G4-CFP and RHAU simultaneously. The presence of RHAU peptide might stabilize G4 structure of mRNA of G4-CFP, resulting in inhibition of the amount of G4-CFP expression in E. coli. That would open a potential peptide candidate for many biomedical applications.

Schematic representation of inhibition of protein expression by RHAU and G4 interaction in
MATERIALS - METHODS
Construction of plasmids
G4 sequence was applied to the upstream of cfp gene (Figure 3). DNA encoding for this CFP protein was amplified by PCR using pHT58218, 17 containing cfp gene as the template with a pair of primer: forward primer ON1: 5'-gcgtagatctgttgggtgggtgggtgggtatgg gcgtgagcaagggcgaggagctgttc-3' and reverse primer ON2: 5'-ccatctcgagttacttgtacagctcgtccatgccgagagtg-3' (IDT, Singapore). This PCR product was then cloned into treated pET-Duet1 (containing two T7 promoters) at BglII/XhoI (New England Biolabs, United Kingdom) sites, resulting in plasmid pD21 (consisting of G4 structure next cfp gene).
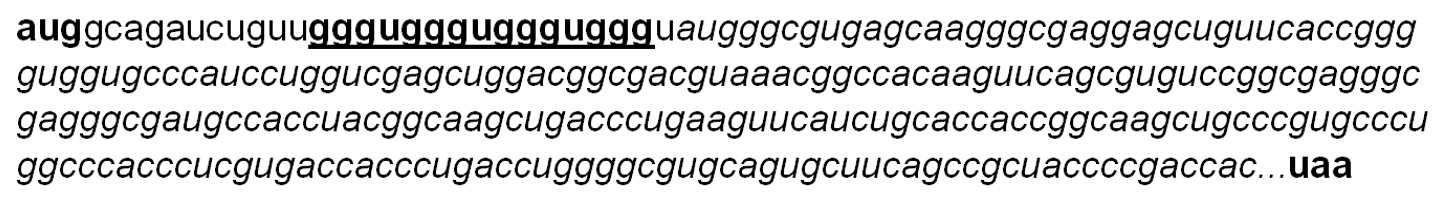
mRNA sequence of
Plasmids pD64 (expressing both G4-CFP and RHAU) and pD65 (expressing both G4-CFP and ΔRHAU-without RSM motif) were generated by cloning DHX36 and ΔDHX36 to the second multicloning site of pD21, respectively. DNAs encoding for RHAU and ΔRHAU were amplified by PCR using RHAU as the template with ON3/ON5 (ON3: 5'-gcgtggatccgtccatgcatcccgggcacctgaaag-3', ON5: 5'-gtgtaagcttctagccgctttttttcttttg-3') and ON4/ON5 (ON4: 5'-gtgtggatccgaaacaggggcagaagaacaag-3') (IDT, Singapore), respectively. PCR products (DHX36 and ΔDHX36) were then cloned into treated pD21 at Bam HI/Hind III (New England Biolabs, UK) sites, resulting in pD64 and pD65, respectively.
Co-expression of protein in E. coli
The plasmid pET-Duet1 containing two T7 promoters that allow expressing two proteins simultaneously. Plasmids pD21 (g4-cfp), pD64 (g4-cfp and DHX36) and pD65 (g4-cfp and ΔDHX36) were transformed into the host of E. coli strain Rossetta (DE3) plysS. The bacteria were cultured in LB medium containing ampicillin at 37C, 200 rpm. When reaching an OD600 of 0.6, IPTG (Sigma Aldrich, Singapore) was added to a final concentration of 0.3 mM. Then the cells were incubated overnight at 16C, 250 rpm before being harvested.
Evaluation of CFP expression by spectrophotometry and statistical analysis
The cells were harvested at OD600 of 1.2. The pellets were re-suspended into the bugBuster protein extraction reagent (Merck, Singapore) plus bezonase nuclease to degrade DNA and RNA. The insoluble debris was removed by centrifugation at 20,000 rpm, 4C. The soluble fraction of proteins was evaluated by the spectrophotometry at the excitation wavelength of 410 nm and emission wavelength of 475 nm. Statistical analysis for the determination of p-value between protein expression levels was calculated in excel.
Evaluation of protein expression by SDS-PAGE and statistical analysis
The pellets of cells were re-suspended into lysis buffer (Tris-HCL 20 mM, sucrose 15%, pH7.4). The soluble fractions were added into loading dye buffer (2% SDS, 100 mM DTT, 10% glycerol, 50 mM Tris-HCL and 0.1% bromophenol blue dye pH 6.8). The samples were then heated at 95ºC in 5 minutes. The insoluble debris was removed by centrifugation at 13,000 rpm in 5 minutes. The soluble fraction of protein was applied to the denaturing polyacryamid gels for electrophoresis. The gels were then visualized with coomassie brilliant blue. The level of protein expression was analyzed by the AlphaEaseFC software. Statistical analysis for the determination of p-value between protein expression levels was calculated in excel.
RESULTS
Construction of plasmid, protein expression
DNA sequence of genes in plasmids pD21, pD64 and pD65 have been confirmed by DNA sequencing (1 BASE, Singapore). The plasmids pD64 or pD65 containing two T7 promoters allowed expressing simultaneously G4-CFP and RHAU or G4-CFP and ΔRHAU under IPTG inducer (Figure 4). All the proteins were expressed in E. coli under IPTG regulation.

Construction of plasmids pD21 (containing G4-CFP alone), pD64 (containing G4-CFP and RHAU) and pD65 (containing G4-CFP and ΔRHAU).
Evaluation of CFP expression by spectrophotometry
The level of CFP protein expression in E. coli was evaluated by spectrophotometry. The plasmid pD21 only bearing g4-cfp was expressed in E. coli under IPTG inducer. The fluorescent intensity of pD21 was observed around 490 a.u and the fluorescent intensity of pD64 and pD65 were observed around 156 a.u and 386 a.u, respectively (Figure 5). Statistical analysis also showed all p-values were <0.05 that determined there was not a significant difference between the means of two group samples of protein expression. The yield of G4-CFP expression of pD65 was 1.2 times lower than that of pD21. It would explain that two proteins (G4-CFP and ΔRHAU) were expressed simultaneously; therefore, ΔRHAU affects the yield of G4-CFP expression of pD65. Interestingly, the yield of G4-CFP expression of pD64 in the presence of RHAU was around 2-fold and 3-fold lower than that of G4-CFP expression of pD65 and pD21, respectively. The presence of RHAU peptide might stabilize G4 structure of mRNA of G4-CFP, resulting in inhibition of yield of G4-CFP expression in E. coli. In contrast, ΔRHAU (without G4 binding domain) does not bind G4 in mRNA of G4-CFP, therefore ΔRHAU was the minor effect of the yield of G4-CFP expression.

Evaluation of CFP expression by spectrophotometry. G4-CFP expression of pD21 was observed around 490 ±6.5% a.u, G4-CFP expression of pD64 and pD65 were observed at 156 ±10% a.u and 386 ±8.2% a.u, respectively.
Evaluation of protein expression by SDS-PAGE
The expression of G4-CFP, RHAU and ΔRHAU were visualized by SDS-PAGE (Figure 6). The result showed both G4-CFP and RHAU of pD64 and both G4-CFP and ΔRHAU of pD65 were observed on the SDS-PAGE that clarified two T7 promoters of pET-Duet1 can control separately the expression of two proteins in E. coli under IPTG inducer. Data also showed a difference in expression level of RHAU (pD64) and ΔRHAU (pD65) was negligible. Therefore, RHAU specific motif (RSM) of RHAU was not affected by the expression of RHAU. Interestingly, the analysis of intensity by the AlphaEaseFC software showed the level of G4-CFP expression of pD64 was 2 times lower compared to the level of G4-CFP expression of pD65. That is in line with the evaluation of G4-CFP expression of pD64 and pD65 by spectrophotometry. The expression of RHAU peptide of pD64 might selectively bind and stabilize the G4 structure of mRNA of G4-CFP, resulting in inhibition of the translational process of ribosomes to mRNA of G4-CFP.

A) Analysis of protein expression by SDS-PAGE. The samples of protein ladder (M), pD64 before (-), and after inducing IPTG (+), pD65 before (-) and after inducing IPTG (+) were applied to the denaturing polyacrylamid gel for electrophoresis. CFP (28 kDa), RHAU (18.5 kDa) and ΔRHAU (16.5 kDa) were all visualized in the gel by coomassive briliant blue solution. B) The relative integrated intensity value of proteins (after addition of IPTG) were analyzed by the AlphaEaseFC programe: integrated density value (IDV) of pD64 (G-CFP around 3557 ± 10%IDV, RHAU around 3392 ± 8.8% IDV), pD65 (G-CFP around 7134±6.7% IDV, ΔRHAU around 3041±11% IDV).
DISCUSSION
Formation of G4 plays a role in many biological processes such as replication, transcription, translation and telomeric maintenance. The stabilization of G4 structure by peptide has recently emerged as a potential approach in the regulation of protein expression. RHAU peptide can selectively bind and stabilize G4 structure via electrostatic interactions between negative charge of phosphate groups and 3 positive charges of amino acids. That allows RHAU peptide to be a potential candidate for the study of stabilization of G4 in many biological processes. Although RSM (16 aa) can be sufficient for specific binding to G4, a lower binding affinity at micromolar range that prevents to study of G4 and RSM interaction in biology. The length of the peptide significantly influences the affinity of binding. The RHAU peptide with 140 aa (consisting of RSM) which can selectively bind G4 at nanomolar range15. The RHAU peptide can selectively bind and stabilize both DNA and RNA G4s that have been studied in the development of the fluorescent probes specific for G4 topologies and ribonuclease for programmable RNA cleavage. Herein, the RHAU (140 aa) selectively binds RNA G4 of mRNA that may prevent the ribosomes from sliding over mRNA. Using vector pET-Duet1 containing two T7 promoters at different regions allows this vector to control the over-expression of two proteins separately. Expression of a single protein being cloned into this vector often yields higher level than co-expression. That explains the expression of a single G4-CFP protein is observed at slightly higher level than co-expression of G4-CFP and ΔRHAU. Interestingly, the level of G4-CFP expression was significantly decreased in the presence of RHAU compared to the level of a single G4-CFP expression and G4-CFP expression in the presence of ΔRHAU. That explains the presence of RHAU may inhibit protein expression of G4-CFP by the interaction of G4 (mRNA of CFP) and RHAU. In contrast, ΔRHAU without RSM can not recognize and bind to G4 structure that does not affect the translational process of ribosomes to G4-CFP. The inhibition of protein expression by G4-stabilized RHAU will be well characterized in the vector containing two different promoters.
CONCLUSIONS
Stabilization of G4 structure by RHAU peptide inhibited expression of G4-CFP in E. coli. G4-CFP and RHAU or G4-CFP and ΔRHAU were able to separately express in E. coli under IPTG inducer. The yield of G4-CFP expression of pD64 in the presence of RHAU was around 2-fold and 3-fold lower than that of G4-CFP expression of pD65 and pD21, respectively. RHAU peptide might selectively bind and stabilize G4 structure of mRNA of G4-CFP, resulting in inhibition of G4-CFP expression. The G4 and RHAU peptide interaction would provide a promising approach for inhibition of unexpectable protein expression in cells.
Abbreviations
CFP: Cyan Fluorescent Protein
G4: G-quadruplex
RHAU: RNA Helicase associated with AU-rich elements
RSM: RHAU Specific Motif
UTR: Untranslated Region
Competing Interests
There is no conflict of interest.
AUTHORS’ CONTRIBUTIONS
T.T.T.T. performed experiments under the supervision of D.T.D. All authors designed experiments, analyzed data. T.T.T.T and D.T.D wrote the paper.
ACKNOWLEDGMENTS
This research is funded by Vietnam National Foundation for Science and Technology Development (NAFOSTED) under grant number 108.02-2017.305.

